Create HDF5 content from a schema#
The hierarchical content of an HDF5 file can be represented as a Python dict instance.
The function silx.io.dictdump.dicttonx() has the following convention for keys and values
to represent HDF5 concepts:
Schema key |
Value type |
HDF5 meaning |
|---|---|---|
|
||
|
||
|
|
|
starting with |
|
|
starting with |
|
|
starting with |
|
|
starting with |
|
The silx.io.dictdump.dicttoh5() function does not parse special key characters or value schema’s
but requires explicit data types for the different types of HDF5 concepts:
Schema key |
Value type |
HDF5 meaning |
|---|---|---|
|
||
|
||
|
|
|
|
||
|
||
|
||
|
Common Usage#
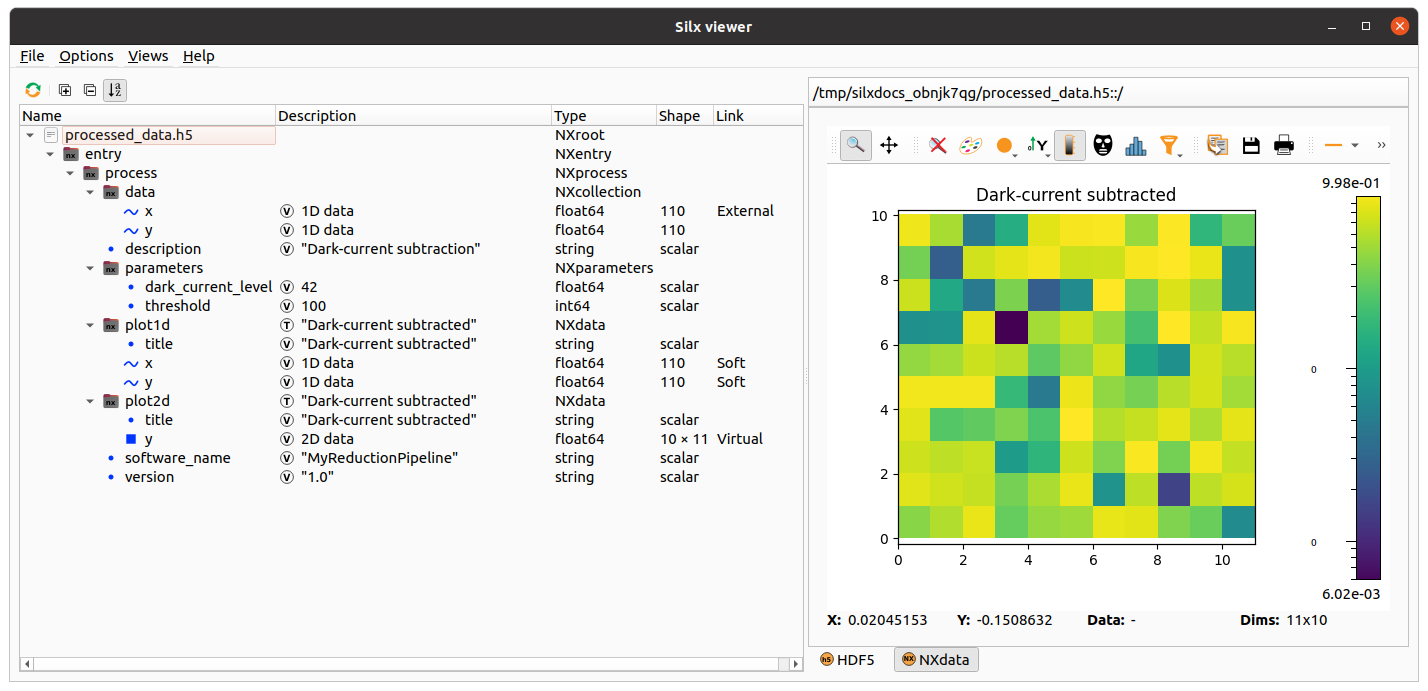
This example
uses a schema describing groups, datasets, attributes, soft links, external links and virtual datasets.
x = numpy.arange(110) / 50
y = numpy.random.uniform(size=110)
data = {
"@NX_class": "NXroot", # HDF5 attribute
"@default": "entry",
"entry": {
"@NX_class": "NXentry",
"@default": "process",
"process": {
"@NX_class": "NXprocess",
"@default": "plot2d",
"description": "Dark-current subtraction",
"software_name": "MyReductionPipeline",
"version": "1.0",
"parameters": {
"@NX_class": "NXparameters",
"dark_current_level": 42.0,
"threshold": 100,
},
"data": {
"@NX_class": "NXcollection",
">x": "./raw_data.h5::/1.1/instrument/positioners/samy", # HDF5 external link
"y": y,
},
"plot1d": {
">y": "../data/y", # HDF5 soft link
">x": "../data/x",
"@signal": "y",
"@axes": "x",
"@NX_class": "NXdata",
"title": "Dark-current subtracted",
},
"plot2d": {
">y": { # HDF5 virtual dataset
"dictdump_schema": "vds_v1",
"shape": (10, 11),
"dtype": float,
"sources": [
{"data_path": "../data/y", "shape": (110,), "dtype": float},
],
},
"@signal": "y",
"@NX_class": "NXdata",
"title": "Dark-current subtracted",
},
},
},
}
raw_filename = os.path.join(tmpdir, "raw_data.h5")
processed_filename = os.path.join(tmpdir, "processed_data.h5")
with h5py.File(processed_filename, "a") as h5file:
dicttonx(
treedict=data,
h5file=h5file,
h5path="/",
update_mode="replace",
add_nx_class=True,
)
with h5py.File(raw_filename, "w") as h5file:
h5file["/1.1/instrument/positioners/samy"] = x
Attributes#
Attributes of groups and datasets can be defined with a key “<item-name>@<attr-name>” or alternatively for groups “@<attr-name>”. In this example we mix both notations:
data = {
"@NX_class": "NXroot",
"entry@NX_class": "NXentry",
"entry": {
"distance": [0, 1, 2],
"distance@units": "mm",
},
}
Soft and External Links#
The target of a soft link is a group or dataset in the same file than the link itself. The target of an external link is a group or dataset in another file than the link itself.
In the example we used this soft link
"data": {
"y": y,
},
"plot1d": {
">y": "../data/y",
}
There are all equivalent ways of defining the same soft link
"plot1d": {">y": ".::/entry/process/data/y"}
"plot1d": {">y": "processed_data.h5::/entry/process/data/y"}
"plot1d": {">y": "/tmp/processed_data.h5::/entry/process/data/y"}
Note
When using “.” as file name it means “the same file as the link itself”. The soft link is always created relative to the link when possible.
Instead of a str the value can also be an instance of silx.io.url.DataUrl. Internally every string
is converted to a DataUrl instance so any format supported by DataUrl can be used.
When the file path in the URL refers to another file, an external link is created. In the example we used
"data": {">x": "./raw_data.h5::/1.1/instrument/positioners/samy"}
Note
The file name of the external link is always converted to a file name relative to the link. The data path portion of the URL must always be absolute in the case.
Virtual Datasets#
Virtual datasets allow merging, slicing and reshaping other datasets in the same file or external files.
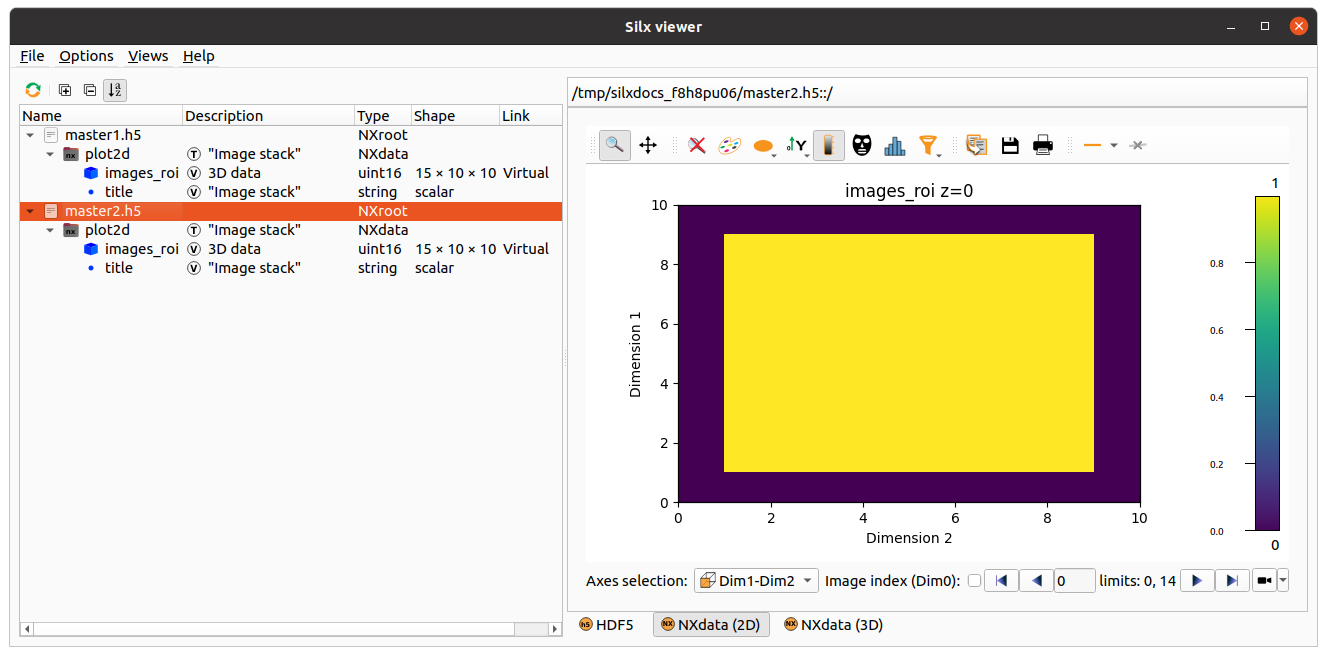
This example uses a list of URL’s to be stacked
in one 3D dataset while selecting an image ROI of [20:30,40:50]:
">images_roi": [
"data0.h5?path=/group/dataset&slice=:,20:30,40:50",
"data1.h5?path=/group/dataset&slice=:,20:30,40:50",
"data2.h5?path=/group/dataset&slice=:,20:30,40:50"
]
Warning
When defining a virtual dataset with a list of URL’s, the source files will be opened and inspected. In addition there is no flexibility in the way the sources are merged together. See Merging URL’s for details on how data is merged (preserve shape vs. stack vs. concatenate behavior).
Here is an equivalent schema that does not open the source files but requires all sources to have the same shape and dtype:
">images_roi": {
"dictdump_schema": "vds_urls_v1",
"source_shape": (5, 50, 60),
"source_dtype": "uint16",
"sources": [
"data0.h5?path=/group/dataset&slice=:,20:30,40:50",
"data1.h5?path=/group/dataset&slice=:,20:30,40:50",
"data2.h5?path=/group/dataset&slice=:,20:30,40:50"
],
}
Here is an equivalent schema that does not open the source files and allows defining the way the sources are merged together:
">images_roi": {
"dictdump_schema": "vds_v1",
"dtype": "uint16",
"shape": (15, 10, 10),
"sources": [
{
"data_path": "/group/dataset",
"dtype": "uint16",
"file_path": "data0.h5",
"shape": (5, 50, 60),
"source_index": (
slice(None, None, None),
slice(20, 30, None),
slice(40, 50, None)
),
"target_index": slice(0, 5, None)
},
{
"data_path": "/group/dataset",
"dtype": "uint16",
"file_path": "data1.h5",
"shape": (5, 50, 60),
"source_index": (
slice(None, None, None),
slice(20, 30, None),
slice(40, 50, None)
),
"target_index": slice(5, 10, None)
},
{
"data_path": "/group/dataset",
"dtype": "uint16",
"file_path": "data2.h5",
"shape": (5, 50, 60),
"source_index": (
slice(None, None, None),
slice(20, 30, None),
slice(40, 50, None)
),
"target_index": slice(10, 15, None)
}
]
}
External Binary Data#
External binary data can be concatenated as a dataset.
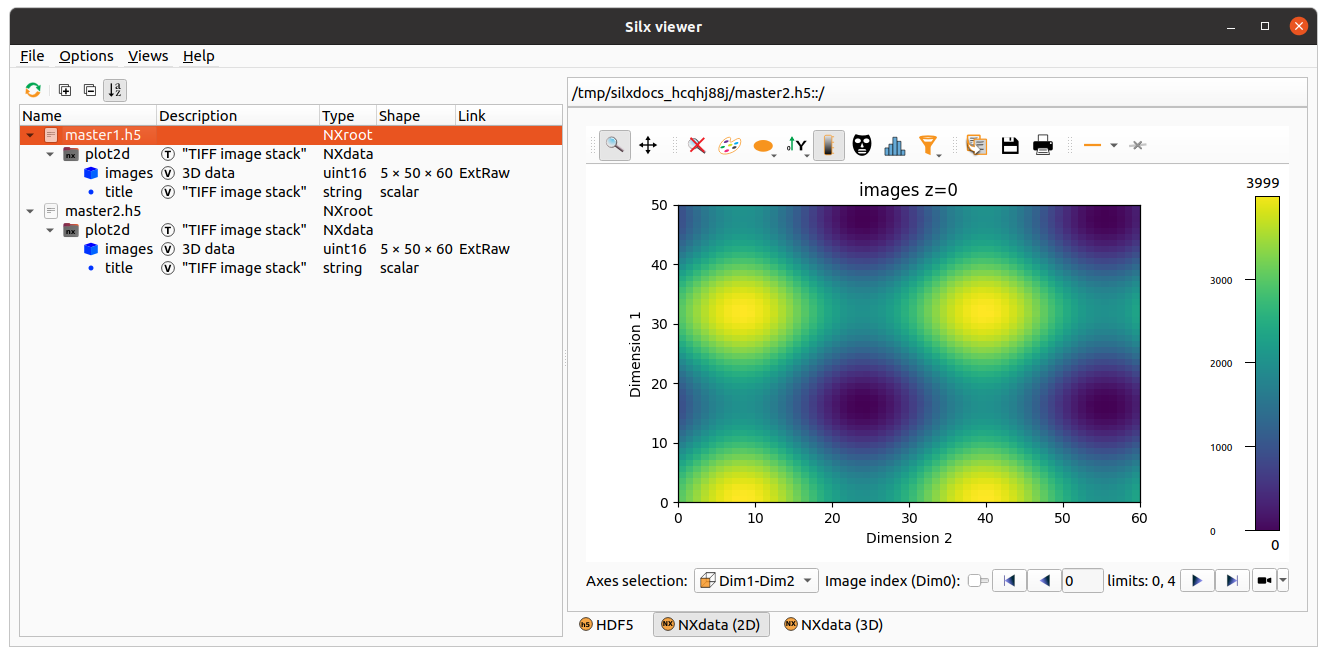
This example uses a list of TIFF files to be concatenated
in one 3D dataset:
">images": ["data0.tiff", "data1.tiff", "data2.tiff", "data4.tiff", "data5.tiff"]
Warning
When defining an external binary dataset with a list of filenames, the source files will be opened and inspected. In addition the HDF5 dataset will store absolute file names so moving the data will break the link. See Merging URL’s for details on how data is merged (preserve shape vs. stack vs. concatenate behavior).
Here is an equivalent schema that can be used for any binary data which is contiguous and uncompressed:
">images": {
"dictdump_schema": "external_binary_link_v1",
"dtype": numpy.uint16,
"shape": (5, 50, 60),
"sources": [
{"file_path": "data0.tiff", "offset": 196, "size": 6000},
{"file_path": "data1.tiff", "offset": 196, "size": 6000},
{"file_path": "data2.tiff", "offset": 196, "size": 6000},
{"file_path": "data3.tiff", "offset": 196, "size": 6000},
{"file_path": "data4.tiff", "offset": 196, "size": 6000},
],
}
Merging URL’s#
One or several URL’s can be merged in a single virtual dataset or external binary dataset.
When providing a single URL, as a string (not a list with one element), the merged dataset has the same shape as the single source.
When providing a list of URL’s, even when the list has only one element, the sources are stacked
when the source rank ndim<3 and concatenated when ndim>=3.
Examples for Nt targets:
target
shape=(): VDS shape(Nt,)target
shape=(N0,): VDS shape(Nt,N0)target
shape=(N0,N1): VDS shape(Nt,N0,N1)target
shape=(N0,N1,N2): VDS shape(Nt*N0,N1,N2)target
shape=(N0,N1,N2,N3): VDS shape(Nt*N0,N1,N2,N3)target
shape=(N0,N1,N2,N3,N4): VDS shape(Nt*N0,N1,N2,N3,N4)…
Warning
Since the sources are merged in a single dataset their shapes must be consistent.
